

PrivaSeq is a toolbase for quantification and analysis of the individual characterizing information leakage, which can be used to link phenotype datasets to genotype datasets and reveal sensitive information in linking attacks.
For technical details please refer to: "Quantification of private information leakage from phenotype-genotype data: linking attacks", Nature Methods, 2016.
The motivation for analysis of linking attacks is motivated by the recent surge of high dimensional phenotyping datasets, which are served with personal information after being "anonymized".
It is becoming clear that we need to proactively evaluate the risks associated with how well an adversary can link the phenotype datasets to the genotype datasets and to other sensitive information.
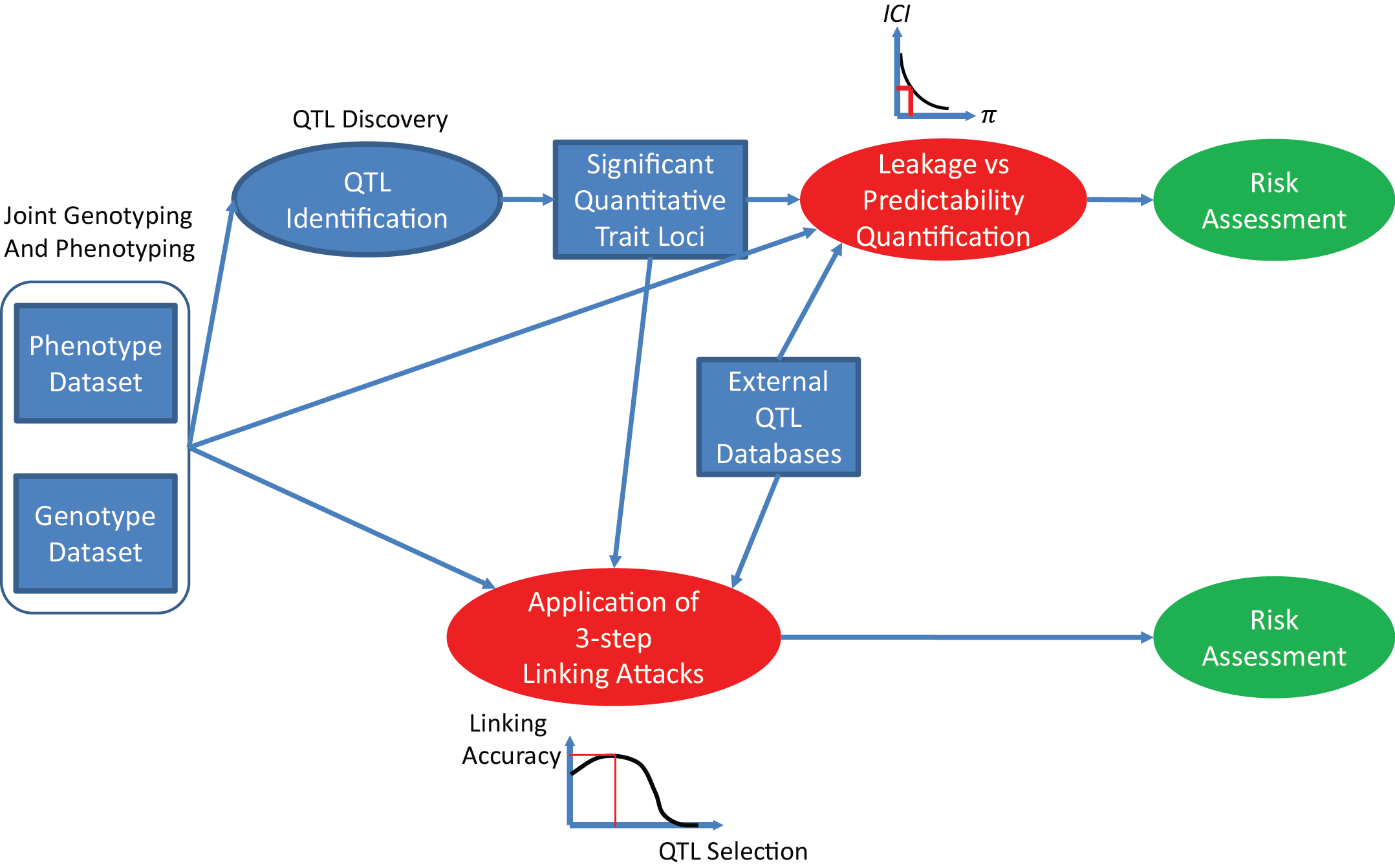
PrivaSeq provides several tools that can be utilized for estimating the information in the phenotype datasets that can be used to link them to genotype datasets. These links are mediated through quantitative trait loci (QTL) datasets, which enable prediction of genotypes from phenotypic information.
The quantifications can be used to direct the data publishing mechanisms to identify the primary sources genotypic information "leakage" in the phenotype datasets and control how much information leaks. These can be used for risk assessment.
We are in the process of building the github repository. In the meantime, you can download the leakage quantification and vulnerable fraction computation code from here. The directory contains a README.txt with file formats that the code uses and an example for running the analysis in the manuscript. Currently, only the leakage from eQTL datasets are considered. The code can evaluate 3 sources of leakage:
We are working on extending the code to easily handle other data formats and other types of QTLs so that the leakage estimates can be computed for other data types.